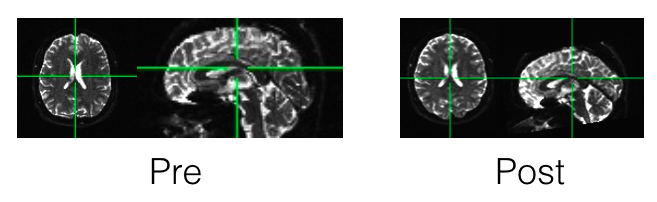
If you’re reading this post either you’re a loyal fan (yay!) or you’ve run Freesurfer (possibly in parallel) and processed your diffusion data in (hopefully TORTOISE) and then Tracula (also possibly in parallel) and you’re wondering where do you go from here. I’m glad you’re here, that’s probably me talking to myself because I read my own blog to remind myself how to do things from past analyses!
So you’ve got your Freesurfer folder and your Tracula folder. The first step is to export all of your data into text files (one per tract is easiest) for all of your subjects. I tend to use a quick bash script to do this, relying on the Freesurfer/Tracula tools tractstats2table to do the hard work (rename Subject* to the prefix of your subjects):
#!/bin/bash
cd /path/to/tracula/folder
mkdir output
for aTract in fmajor_PP_avg33_mni_bbr \
fminor_PP_avg33_mni_bbr lh.atr_PP_avg33_mni_bbr \
lh.cab_PP_avg33_mni_bbr lh.ccg_PP_avg33_mni_bbr \
lh.cst_AS_avg33_mni_bbr lh.ilf_AS_avg33_mni_bbr \
lh.slfp_PP_avg33_mni_bbr lh.slft_PP_avg33_mni_bbr \
lh.unc_AS_avg33_mni_bbr rh.atr_PP_avg33_mni_bbr \
rh.cab_PP_avg33_mni_bbr rh.ccg_PP_avg33_mni_bbr \
rh.cst_AS_avg33_mni_bbr rh.ilf_AS_avg33_mni_bbr \
rh.slfp_PP_avg33_mni_bbr rh.slft_PP_avg33_mni_bbr \
rh.unc_AS_avg33_mni_bbr
do
echo $aTract
tractstats2table --inputs Subject*/dpath/${aTract}/pathstats.overall.txt \
--overall -t output/${aTract}.txt -v
done
Now that you’ve exported all the numbers to text files, the next step is often to combine all of those outputs into a single file for data analysis. You might have already had the great idea of using “cat” to combine the files via the command line, and then you’d be disappointed that it’s not that easy because the header of the first column (the one with your subject numbers) is the tract name, but there’s no dedicated column for the tract name (at least not in Freesurfer 6.0).
So here’s some handy R code:
library(tidyverse) #if you don't have tidyverse, you should.
setwd('/path/to/tracula/folder/output') #folder with text files from bash script
allFiles <- list.files(pattern='.txt') #get a list of all the text files
#functions make the world go 'round:
#this one reads in each file, makes a column with the tract name
#and renames the first column to "Filename"
readDTI <- function(x) {
tmp <- read.table(x, header=T)
tmp$tract <- names(tmp)[1] #get the tract name and make it a column
names(tmp)[1] <- 'Filename' #now make the first column a useful name
tmp$Filename <- as.character(tmp$Filename) #useful if you want to manipulate the header later
return(tmp)
}
#this will run your function on all the text files you found with list.files()
alltracts <- lapply(allFiles, readDTI) #lapply returns a list
#this will return a single data.frame object with all the tracts
tracula_data <- do.call("rbind", alltracts)
#this writes the whole thing to a file on your computer
write.table(tracula_data, "MyTraculaData.dat", sep="\t", row.names=F, col.names=T, quote=F)
And just like that, you have a single tab-delimited text filled with all your data for 18 tracts (9 per hemisphere) for all of your subjects) for reading into your favorite stats program, or sending to co-authors.