These instructions are for an older version of TORTOISE, if you would like to read new instructions checkout the updated tutorial HERE!
There are many options when deciding how to process Diffusion Weighted Images (DWI) and turn them into Diffusion Tensor Images (DTI). I’ve written before about preprocessing in FSL (Part 1, Part 2) and AFNI (Part 1, Part 2), highlighting that most recommend doing some kind of eddy current “correction”, rotating the b-vectors to adjust for motion in the scanner, and often registering the data to an individuals brain. But the capabilities are often spanning different software packages or not completely handled by the tools forcing you to write your own code to handle bvec rotation or coregistration. Don’t get me wrong, I usually enjoy this! But there is something to be said about using a canned software package that takes care of all of the preprocessing steps!
Enter TORTOISE (Tolerably Obsessive Registration and Tensor Optimization Indolent Software Ensemble). TORTOISE can handle all of the preprocessing of your DWI data with full support of eddy and motion correction, BSpline correction, rotation of b-vectors, and co-registration to a structural image. TORTOISE can also fit the tensors using both a linear fit (similar to FSL), nonlinear fit, and weighted non-linear methods (e.g. RESTORE and iRESTORE). In this post I’ll go over the preprocessing of DWI images (DIFF_PREP) and next time I’ll go into more depth on the calculations via DIFF_CALC. The recommendations I make are based on both the written manual (accessible on their wiki) and my own personal experience using TORTOISE. I highly recommend using TORTOISE, which requires no additional software. If you have a copy of IDL on your computer (or are willing to buy a copy), TORTOISE will run faster by parallelizing many of it’s operations.
Preprocessing DWI Data with DIFF_PREP
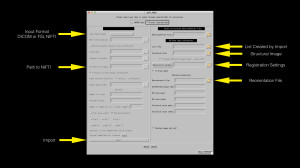
The first thing to do is get your data into TORTOISE. You can import a variety of data formats (Phillips PAR/REC, DICOM, NIFTI, and Bruker). I highly recommend importing from either DICOM or FSL NIFTI. If you choose DICOM, you will need to supply TORTOISE with your gradient file, as it will not read the directions from your DICOM headers. If you choose FSL NIFTI, TORTOISE will read your bvec and bval files from text, they just need to have the name (bvec or bval) in the filename and be in the same folder as the NIFTI file. TORTOISE will search sub-folders for these files, so if you have multiple bvec or bval files in your folder or sub-folders, it will give you an error that it found multiple bvec/bval files and you’ll have to tidy up your file structure.
In the image above (click to make bigger), you can see the common settings highlighted with yellow arrows. On the import side, you’ll want to specify your format (FSL NIFTI is my recommendation), the file path to your NIFTI file (note – uncompressed NIFTI only -.nii not .nii.gz – as of this post), and then click import. Using NIFTI files imported from dcm2nii and DIMON have worked well for me. The one advantage to dcm2nii is that it will automatically create the bval and bvec files in the correct format for TORTOISE.
After you import the data, you will see the right side of the window becomes enabled and will automatically file in your List File location. If you have a T2-weighted image with fat suppression, you should supply that as your Structural File. This structural file should be skull stripped and optionally put into AC-PC alignment. If you don’t have a T2-weighted image, you will want to turn off the BSpline correction step. I’ve found that you can use a T1-weighted image for as Structural File so long as you don’t use BSpline correction. If you don’t supply it a Structural Image, TORTOISE appears to make one from the B0 image. The bottom line is that in this particular software – having a T2-weighted image for correction is very helpful.
If you have a high-resolution T1-weighted image that you wish the DWI images to be in alignment with, you can supply a Reorientation File as well. This could be your skull-stripped anatomical image from an AFNI or FSL pipeline in the desired final orientation (e.g. RAI). It could also be a template (e.g. MNI152). It’s worth noting that TORTOISE is using a standard linear (“affine”) transform, which may not be as good as a non-linear transform. But has the advantage of being part of a transform chain that only interpolates your data once!
Next you’re going to click the “Registration param” button to setup the more advanced settings.
 Inside your Registration Settings, I highly recommend you keep most of the defaults. The things that I’ve found reason to change (in TORTOISE version 2.0.1) are the final DWI voxelsize resolution, it defaults to 1.5mm^3, but it may not make sense to go to such a fine grid if you collect at say 3mm^3. The other thing that I’ve changed is the initial upsampling setting, if you have a particularly thick slice DWI image, you might consider changing it to slice_only. In the registration settings is also where you will turn off the BSpline correction if you aren’t using a T2-weighted image with fat suppression for your structural image.
Inside your Registration Settings, I highly recommend you keep most of the defaults. The things that I’ve found reason to change (in TORTOISE version 2.0.1) are the final DWI voxelsize resolution, it defaults to 1.5mm^3, but it may not make sense to go to such a fine grid if you collect at say 3mm^3. The other thing that I’ve changed is the initial upsampling setting, if you have a particularly thick slice DWI image, you might consider changing it to slice_only. In the registration settings is also where you will turn off the BSpline correction if you aren’t using a T2-weighted image with fat suppression for your structural image.
Once you click “Apply”, TORTOISE will start working. It can take a while, so be patient. Some windows will popup to show the registration of each direction to the B0 image and the registration of the B0 to the structural image.
Why use TORTOISE?
If you’re still reading, you might be saying to yourself “Why would I add another software package to my existing pipeline?” Well that’s a fair question and one that I asked repeatedly, until I saw a demonstration by Paul Taylor (author of many of the AFNI tractography programs) showing the difference in tractography with and without TORTOISE processing, the presentation can be seen here, see slide 52.
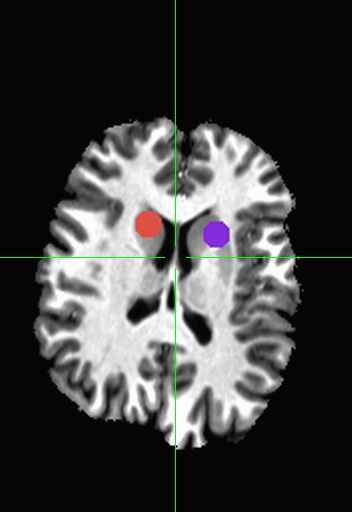
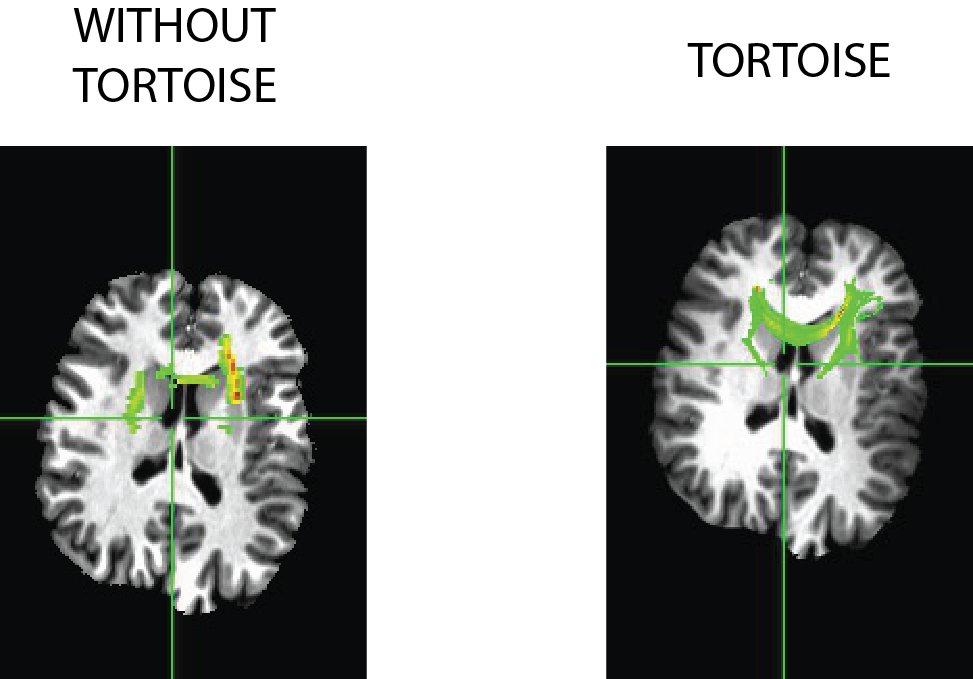
I went ahead and made a quick “off the cuff” comparison of using TORTOISE vs. only doing eddy correction (via FSL’s eddy_correct) tool. This sample with TORTOISE uses all of the default and suggested options listed above, but without a T2-weight structural image (I didn’t have one). TORTOISE rotated the bvecs automatically, but I did not rotate the bvecs on the FSL pipeline, though there was relatively little movement, so I wouldn’t expect the rotation to make a big difference. Tensors were fit via a linear fit method and then deterministic tractography was used (LOGIC: AND) between two frontal ROIs shown below:
There are plenty of fibers that appear in the NON-TORTOISE output, but considerably more connections in the TORTOISE output. It appears that the fibers are partially going through the ventricles, this is mostly an illusion based on the slice and viewing direction, the tract in reality (mostly) goes around the ventricle.
Stay tuned for the tensor fitting segment via TORTOISE (DIFF_CALC). I will also get around to describing how you can use TORTOISE with your AFNI processing pipeline.